Research Staff:
Processing genomic and medical information efficiently and at scale
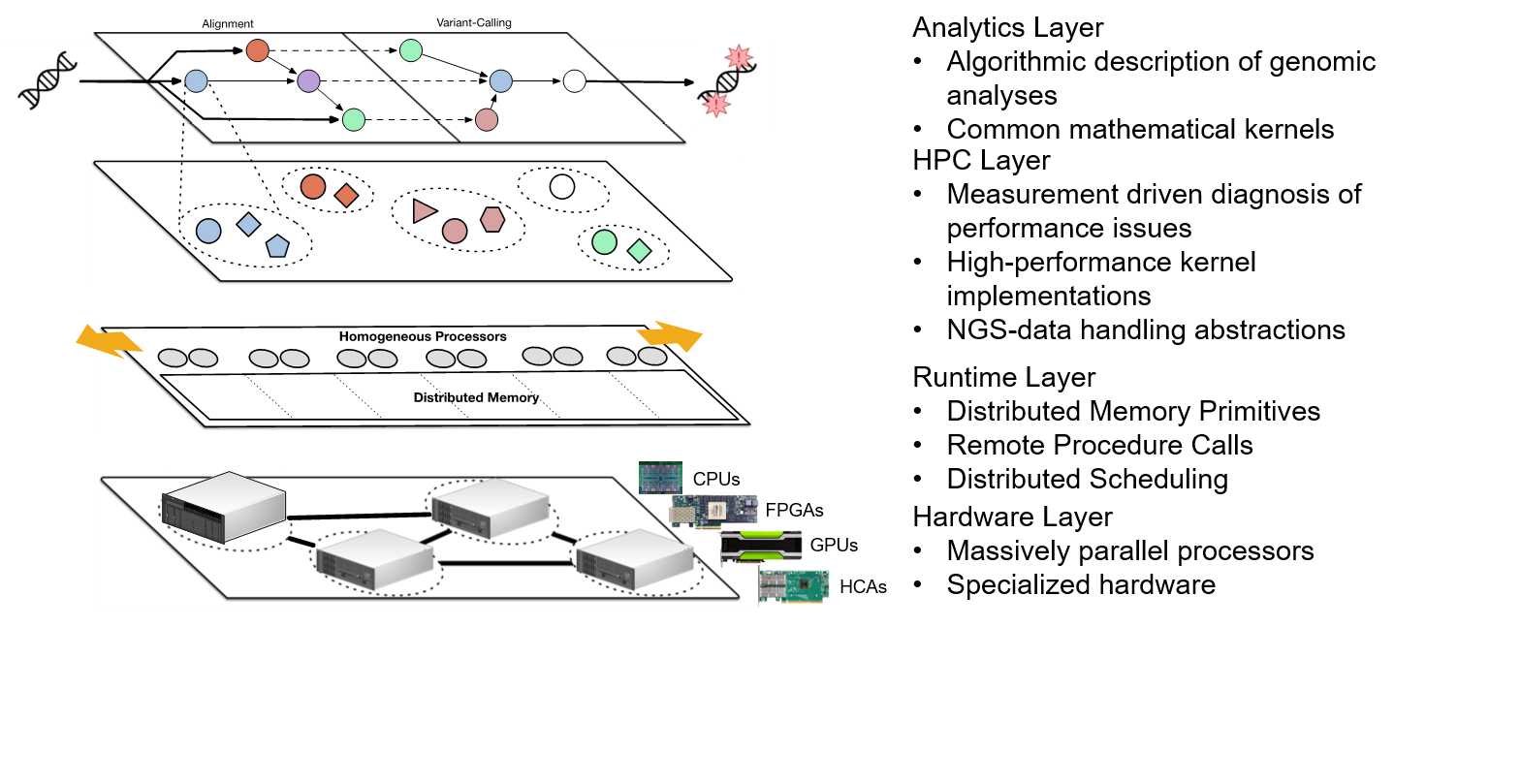
This project consinsts in designing and developing systems to efficiently compute the germline and somatic variants (mutations) from raw sequence data. The work encompasses several key aspects of the CompGen instrument.
(1) Static algorithmic analyses to identify the core computational kernels used in processing genomic data for personalized medicine, which enable a targeted study of performance and accuracy of a vast number of genomic applications and enable the implementation of a system-level framework to accelerate them.
(2) Run-time environments for execution of computational genomics workloads on clusters of heterogeneous processors, which enable instrument users, such as biologists or clinicians, to express their computations at a higher level, without having to worry about the computer system-centric details of their implementations. In particular, the instrument’s run-time system “Symphony” developed by the DEPEND researchers, can transparently decide on scheduling computations based on processor affinity, data locality, and resource contention.
(3) Applying algorithmic approximations that allow for more efficient execution of computational genomics workloads, without any loss in accuracy. In particular, our group designed an approximation algorithm for Levenshtein distance computation in sequence alignment that represents the calculation as operations on an algebraic lattice, like time, thereby allowing for significantly more efficient hardware implementations as compared to traditional methods.
(4) Building custom hardware accelerators for computational genomics. In collaboration with IBM, the DEPEND group has developed accelerators for computationally intensive tasks in human variation detection and genotyping, such as DeBruijn assembly, Alignment, and PairHMM kernels.
FPGA acceleration of Edit-distance & PairHMM using CAPI
By using the coherent accelerator processor interface (CAPI), our team of students and professors, in collaboration with Prof. Deming Chen and his students (Jong Bin Lim) have developed a hardware accelerator to improve the runtime of short-read alignment. The proposed design was implemented on an Altera Stratix V FPGA in an IBM POWER8 system using the CAPI interface for cache coherence across the CPUs and FPGA. The suggested design is 200 times faster than the equivalent C implementation of the kernel running on the host processor and 2.2 times faster for an end-to-end alignment tool for 120-15bp short-read sequences. Currently, we are working on accelerating another gnomic kernel (PairHMM) by building an array of processors, which is based on having many PEs(processing elements) (data-path elements) controlled by schedulers (control flow elements). Each PE-scheduler pair computes a separate instance of the PairHMM kernel in parallel. This will allow us to scale to arbitrary size inputs make the best use of the fast SRAM available to us on the FPGA.